Big implications: Individual variations
It has been shown that certain pharmaceutical agents can be effective in some patients but fail to work for others or lead to serious side effect. Human variability regarding the safety and effectiveness of medications depends on several factors, the most important being the difference in genetic composition between one person and another. Essentially, the varying responses are based on the 0.01% difference in the genetic makeup of individuals.

Heterogeneity: The impact of tumour variations
Cancer cells tend to be genetically unstable. The genetic makeup of cancer cells in patients that have been diagnosed with the same kind of cancer can vary! There may also be significant genomic variance between cancer cells in a particular patient (Yao and Dai, 2014).
This understanding lead to a revolutionary leap into a completely new era of medicine: personalised medicine, also called ‘precision medicine’.
Factoring the genomic profile of a specific cancer into decisions, along with the patient’s environment and lifestyle, allows the medical professional to tailor the therapy in order to be more effective. This also includes careful monitoring of the patient response to a particular treatment and adjusting the treatment if necessary.
Breast cancer, for instance, has been found to be highly heterogeneous, even to a point where the molecular and morphological characteristics contrast between patients. A targeted and patient-specific therapy approach seems more promising than a traditional ‘one size fits all’ approach.
Precise medicine: The application
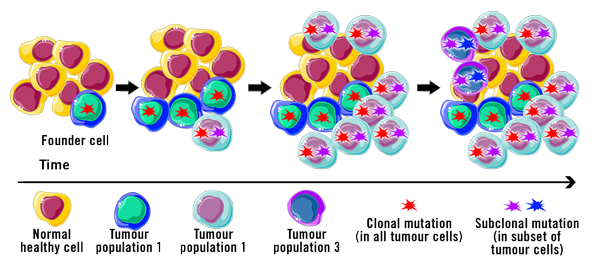
One hallmark of cancer is genomic instability, such as when genes for regulatory pathways involved in DNA repair are disrupted. This can lead to increased replication errors and defects in the mitosis machinery. Consequently, a variable genomic profile occurs in cancer cells after each mitotic division. This constant change could, in turn, reduce the effectiveness of a treatment.

Cancer development
Tumours share several common traits and capabilities. During the development, the tumour cell is subject to genome instability that are involved in the following capabilities of the tumour cell – the hallmarks of cancer:
- Sustaining proliferative signalling
- Evading growth suppressors
- Avoiding immune destruction
- Enabling replicative immortality
- Tumour-promoting inflammation
- Activating invasion and metastasis
- Inducing angiogenesis
- Genome instability and mutation
- Resisting cell death
- Deregulating cellular energetics
(Hanahan and Weinberg, 2011)
A model patient suffers from lung cancer, specifically from bilateral stage IV well-differentiated adenocarcinoma. Genomic profiling of the tumour tissue shows a deletion on exon 19 of EGFR. This results in the expression of an epidermal growth factor receptor (EGFR). However, this EGFR is a constantly active tyrosine kinase receptor that induces cellular proliferation. A specific treatment to inhibit EGFR function leads to tumour remission.
After 24 months of treatment, cancer recurrence has been detected. Analysis of the tumour DNA shows an additional mutation in the EGFR gene called T790M. Due to this mutation, the first therapeutic agent is not effective anymore, and has to be replaced by a second agent that inhibits EGFR again. Two months later, the tumour goes into remission.
Trials with more than 1,000 patients diagnosed with non-small-cell lung carcinoma (NSCLC) showed that genotype-directed treatments resulted in longer survival (Gambardella et al., 2020).
NGS in oncology: Clinical diagnostics tool
For tumour analysis, usually, a tissue sample is obtained through a biopsy. The extracted DNA is then subjected to high-throughput sequencing, and the resulting data provides information that aids the selection of the most effective therapy, and predicts the adverse effects of a treatment, specifically a medication.
Whole exome sequencing (WES) of tumour-derived DNA enables the identification of somatic and germline mutations. Somatic mutations could correspond to tumour-specific genomic aberrations. Specific germline variations can indicate the risk of cancer development in predisposed individuals.
A targeted panel approach analysis of a certain subset of cancer-specific genes. These panels often include between 20 to 150 genes that are specific for certain cancers, such as breast cancer or lung cancer.
Whole exome and targeted gene sequencing is usually done for solid tumour biopsies. However, extraction of solid tumour biopsies is not always feasible and usually highly invasive. Liquid biopsy is an approach that makes use of minimal invasion, and can also be used to profile the whole exome, a larger range of specific key cancer-associated genes, or single specific mutations in a tumour sample.
Comprehensive genomic profiling (CGP), on the other hand, analyses the full coding sequence (CDS) of around 600 cancer-specific genes, as well as promotor regions, miRNA sequences, extra-exonic variants and gene fusions. CGP shows the highest sensitivity due to the hybrid capture-based target enrichment and a coverage greater than 500x. This so called pan-cancer analysis, which aims at examining the alterations across diverse tumour types, is much more comprehensive than panel sequencing and is available for a more attractive price than WES.
NGS in oncology: Liquid Biopsy
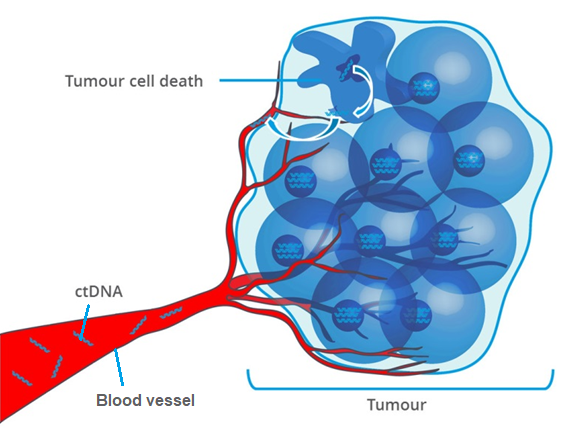
Cell-free DNA (cfDNA) is constantly released into body fluids. Most of the cfDNA originate from cells that undergo cell death, but is also secreted by living cells.
Similarly, cancer cells also release DNA fragments, called circulating tumour DNA (ctDNA) into the bloodstream. ctDNA can be used to genetically profile the tumour as the genetic alterations in ctDNA are identical to those in the tumour source. The analysis includes point mutations, chromosomal rearrangements, epigenetic patterns and copy number variations (CNVs), amongst others (Parikh et al., 2019).
Forerunners and experts for NGS in oncology
The Oncotype DX Genomic Prostate Score (GPS) is a forerunner in the utilisation of genomic testing in cancer treatment. For men diagnosed with prostate cancer, the GPS helps to identify the immediacy of treatment initiation. Thus, with the use of this test, both the probability of metastasis and death resulting from this type of cancer can be quantified. Consequently, if the test results show malignant tumours that are confined to the prostate, the success of the treatment can be continuously monitored (clinical surveillance). If a more aggressive type of cancer is involved, the most favourable medications and solutions for the patient can be studied.
Foundation Medicine is a global provider of diagnostic tests based on CGP for cancer patients at advanced stages. These companion diagnostic tests analyse clinically relevant biomarkers and genomic alterations, and are utilised to determine the applicability of a therapeutic agent to a specific person. Physicians gain insights that potentially optimise cancer treatment for their patients.
The company Oncompass Medicine is one of the first providers of genomic analysis of solid and liquid biopsies in Europe. The company’s high experience and great innovative spirit in the field of diagnostics in oncology was honoured with the Future Unicorn Award in 2021. This award is given to scale-ups that could become the future tech giants of Europe!
Oncompass uses solutions for the simultaneous analysis of nearly 600 cancer-associated genes, whole exome analysis, and analysis of circulating cancer DNA (cfDNA) in liquid biopsies. Their sophisticated analysis software and knowledge database facilitates the selection of the most effective treatments for each patient and continually evaluates the effectiveness of new types of therapies.
Conclusion
Valuable information can be gained from the analysis of genomic data to support a more accurate diagnosis. Therapeutic management can be optimised to promote an improvement in the quality of life. In addition, the molecular profile of each individual tumour can be further used in clinical research. This makes the increased application of genomic testing in the field of oncology a benefit not only for the prevention, detection, and management of cancer, but also to improve health-care for future generations.
NGS is successfully used as as clinical diagnostics tool outside of oncology.
By Valeria Vélez Potes and Dr Andreas Ebertz
Did you like this article on NGS as clinical diagnostics tool in oncology? Then subscribe to our Newsletter and we will keep you informed about our next blog posts. Subscribe to the Eurofins Genomics Newsletter.






11 Comments