Brief overview: Vaccines and the immune system
Conventionally, vaccines contain attenuated viruses that alarm and prepare the immune system against future invasion by pathogens that contain similar characteristics. In the case of whole viruses, scientists either engineer viruses that raise fewer symptoms or use non-functional viruses. In the late 20th century, the research focus shifted to identifying the key molecules on the virus surface that interact with immune cells to activate an immune response. The basic idea of a vaccine is to train the immune system to recognise and fight a pathogen without exposing the host to the full pathogenicity.

The problem: Production of vaccine
Producing conventional vaccines by engineering viruses or viral subunits is a costly and time-consuming process. In case of the flu vaccine, it takes approximately five to eight months from vaccine development to the approval for public use. The limiting factor of the production of subunit or live-attenuated vaccines is the requirement of cell cultures. Scientists have been making efforts to devise fast ways to produce an effective vaccine.
The solution: mRNA vaccine technology?
One of the strategies to speed up the production of effective vaccines is the implementation of mRNA. In 1990, scientists working at the University of Wisconsin provided the first evidence of gene expression mediated by injected RNA in mice muscle.
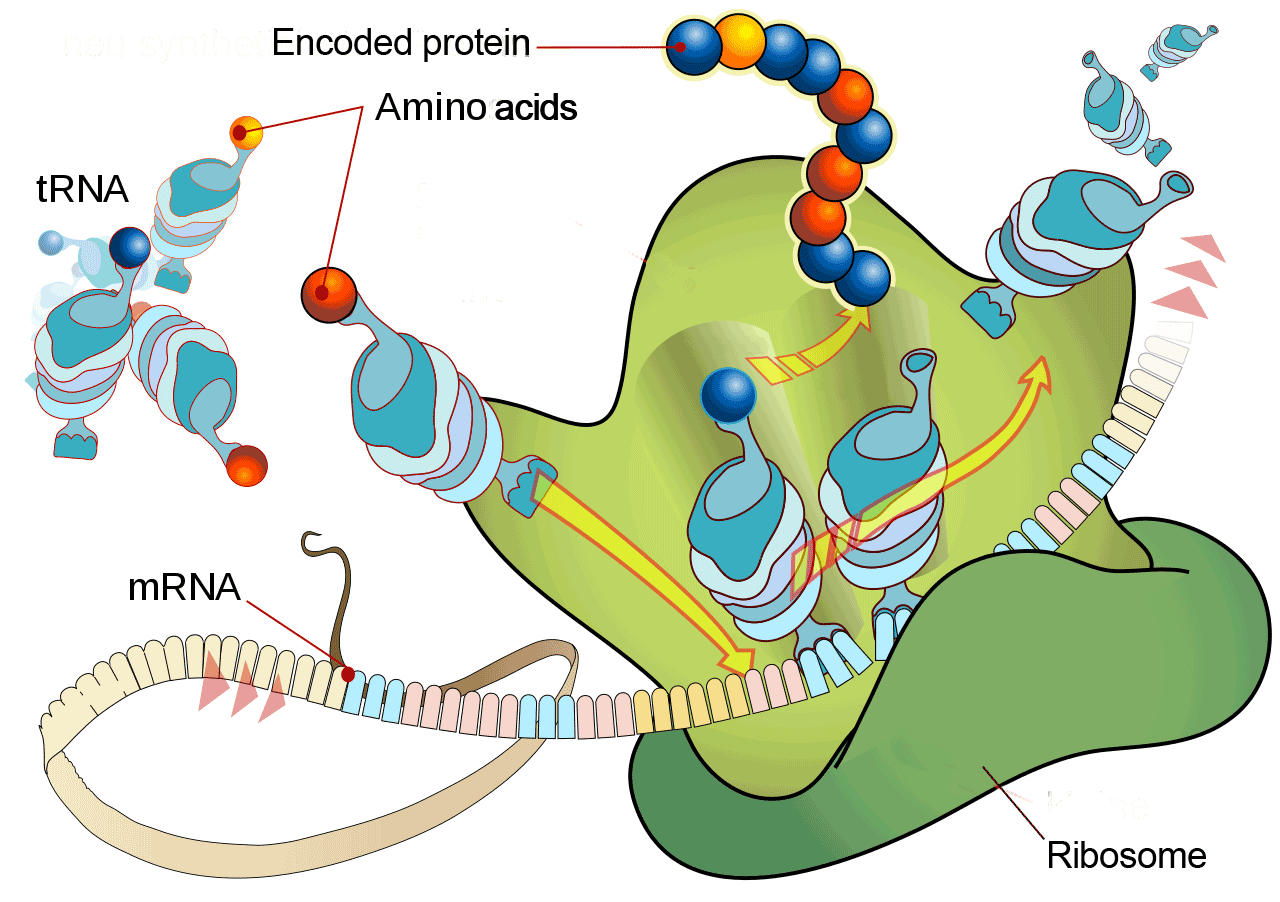
RNA vaccines exploit the gene expression mechanism of the cell. The vaccine consists of mRNA encoding the antigen. Once inside the cell the mRNA is translated into the antigens, which is equivalent to hijacking the cells. The antigens are displayed on the cell surface and are recognized by the immune system (Wolfe et al., 1990).
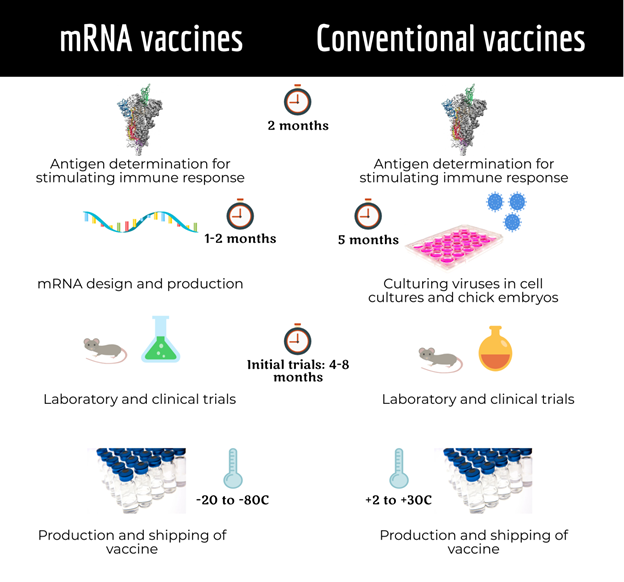
The effective response to a pandemic entails the earlier availability of vaccines. However, the meticulous process of development, trials and approval takes a certain period of time. In that regard, the nature of the mRNA vaccine is highly beneficial as it allows for quicker design and production compared to conventional vaccines.
Hekele et al. have designed a lipid nanoparticle formulated self-replicating RNA vaccine against the H7N9 influenza strain in only 8 days (Hekele et al., 2013). As crises are often drivers of innovation, robust and prompt RNA technologies have emerged in the face of COVID-19 pandemic (Bloom et al., 2020).
mRNA vaccine technology: Three decades in the making
Although, the first successful in-vivo gene therapy that used mRNA was reported in 1990, it took three decades of research until the first mRNA vaccine was available. The main challenges on the development road are delivery and the poor stability of mRNA. The mRNA molecule is highly susceptible to degradation by the immune system.
Katalin Karikó, a renowned Hungarian biochemist, did not give up her efforts. She recognised the potential of mRNA way before the world could. Along with Drew Weissman, she found the secret to prevent activation of the immune response against the intruding mRNA: modification!
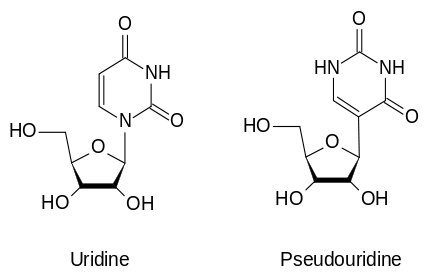
RNA is comprised of the nucleosides Adenosine (A), Guanosine (G), Cytidine (C) and Uridine (U). It has been found that RNA activates the toll-like receptors (TLR) on immune cells. Karikó and Weissman modified the RNA by inserting a naturally occurring modified nucleoside, pseudouridine. The result: this modified RNA transcript does not activate the TLR response and provides a higher translational capacity (Karikó et al., 2005; Karikó et al., 2008). “Karikó and Weissman figured out that if you incorporate modified nucleosides into mRNA, you can kill two birds with one stone,” Norbert Pardi, a colleague at University of Pennsylvania, said.
During these three decades, research on RNA vaccines against cancer, rabies, cytomegalovirus, influenza virus and Zika virus continued in laboratories and clinics (Verbeke, Lentacker, De Smedt, & Dewitte, 2019). Currently, two types of RNAs are used for RNA vaccines.
The types of mRNA vaccine technology
In order to produce an mRNA vaccine, the DNA sequence of the desired gene (coding for the antigen) is transcribed. This mRNA is then modified by adding untranslated regions (UTRs), a 5’ cap and a 3’ polyA tail. This way, the synthetic mRNA mimics the host cell’s mRNA. For the final formulation, various reagents are combined to improve the stability and cellular uptake of the vaccine.
The types of RNAs that are used for RNA vaccines are: non-replicating mRNA and in-vivo self-replicating mRNA.
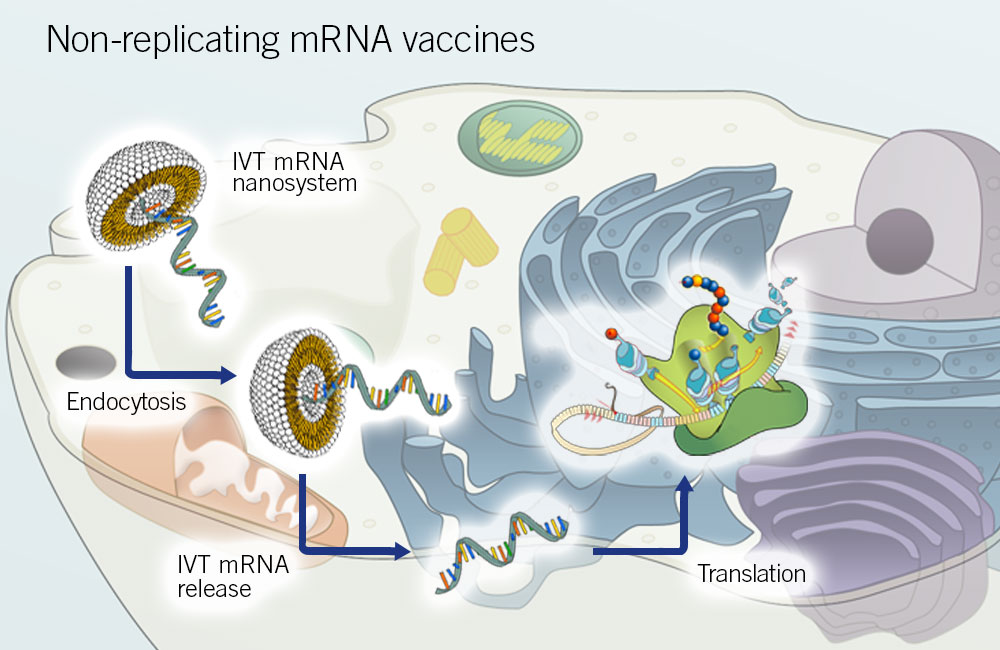
Non-replicating mRNA vaccines
These vaccines consist of a small mRNA containing the antigen-encoding region and flanking UTRs. The mRNA is part of a liposome complex that is taken up by cells. These vaccines are the simplest and most economical ones.
In 2012, the first non-replicating mRNA vaccine was produced against the influenza A virus. The vaccine exhibited a balanced and long-lived protective immune response against the influenza viruses in mice (Petsch et al., 2012).
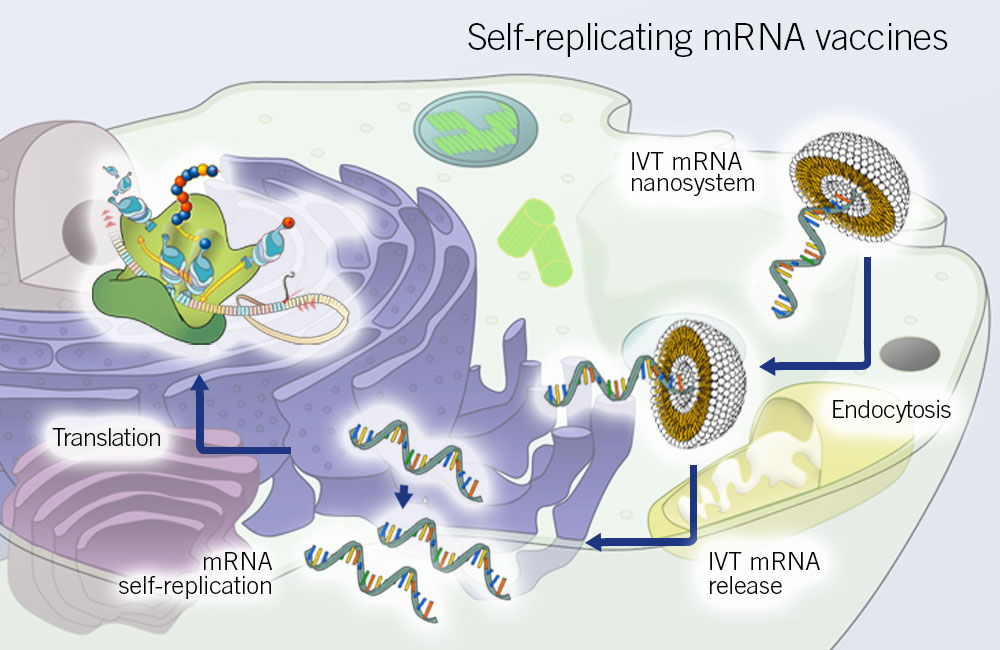
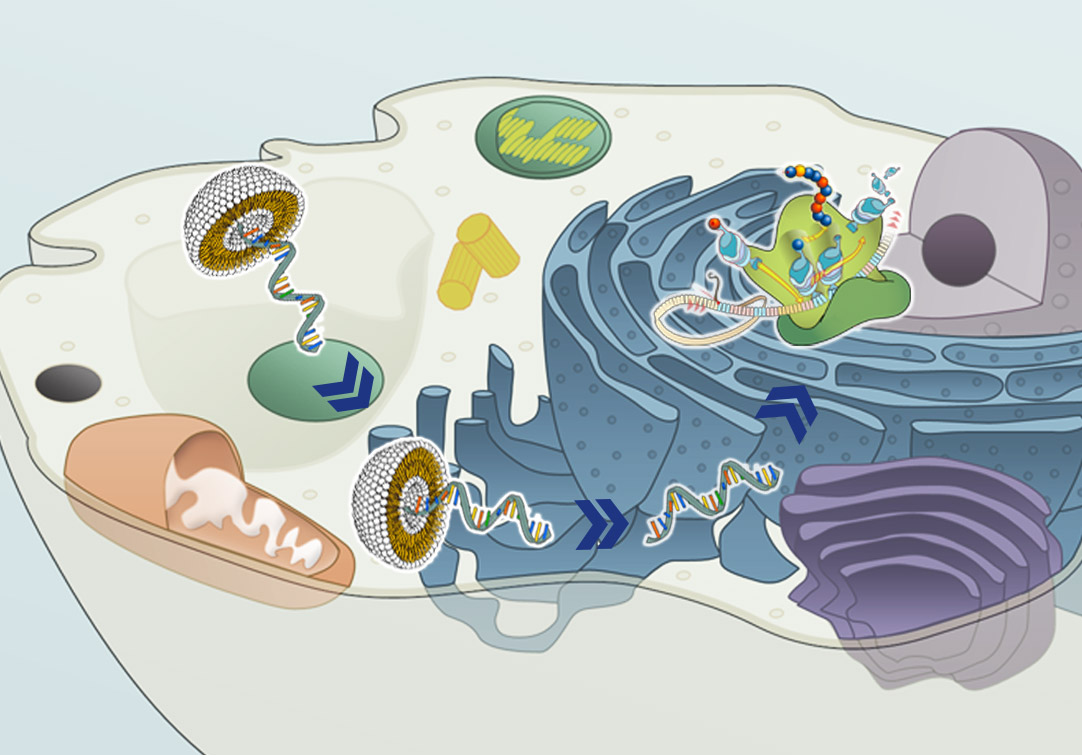
Self-replicating mRNA vaccines
These vaccines consist of an mRNA that encodes the antigen and the viral RNA replication machinery. Currently, the alphavirus genome is used for generating self-replicating mRNA vaccines. In the virus genome, structural proteins-encoding genes are replaced with the antigen-encoding genes, and simultaneously, the RNA replication machinery encoding genes stay intact (Perri et al., 2003).
This type of vaccine ensures a large amount of mRNA production from a very small vaccine dose owing to intra-cellular replication. It triggers a robust immune response. However, the size constraints for self-replicating mRNA vaccines limit their use (Pardi, Hogan, Porter, & Weissman, 2018).
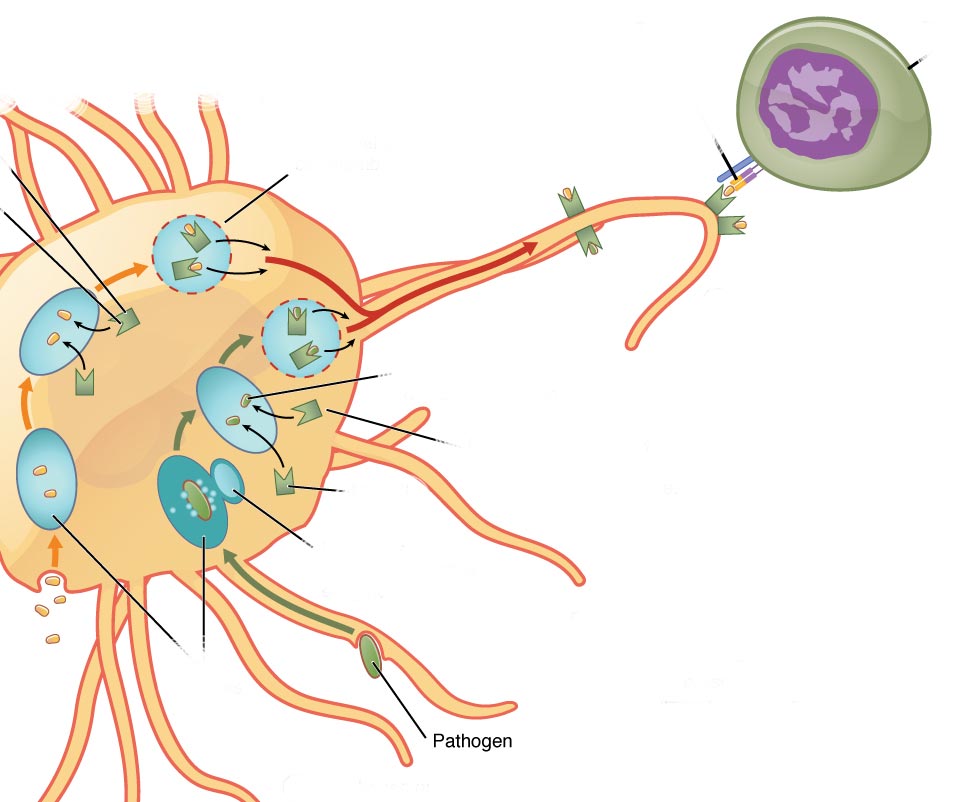
In vitro dendritic cell non-replicating mRNA vaccine
Dendritic cells are a type of immune cells that are specialised in presenting antigens to other immune cells in order to mount an immune response. For this type of RNA vaccination, the host dendritic cells are extracted and transfected with the mRNA transcript. Upon successful transfection, the cells are delivered back to the host.
This type of vaccine has been used extensively in anti-tumour response laboratory and clinical trials (McNamara, Nair, & Holl, 2015). Although, its use has been limited to HIV in infectious disease studies (Pardi et al., 2018).
Pfizer-BioNTech and Moderna launch the first mRNA vaccines
In December 2020, FDA and other regulatory authorities in the UK and Europe approved the use of the two mRNA vaccines by Pfizer-BioNTech and Moderna. Both are non-replicating mRNA vaccines and use modified mRNA transcripts that code for SARS-CoV-2 spike protein. The vaccines show more than 90% efficacy and success rate. It took only 25 days to produce the 1st batch of Moderna’s mRNA vaccine against SARS-CoV-2 (Corbett et al., 2020; Jackson et al., 2020). BioNTech submitted a request for vaccine approval within 200 days after the beginning of the vaccine development program (Dolgin, E. 2020).
Learn more about the genomic make-up of SARS-CoV-2. What proteins are encoded in the SARS-CoV-2 genome? How can this information be used to identify SARS-CoV-2?

Both vaccines are similar in the use of mRNA, mode of delivery (intra-muscular injection) and dosage requirements. They require a booster shot after the primary dose. The interval between the doses for Moderna’s vaccine is 28 days, while for Pfizer-BioNTech’s vaccine is 21 days. However, the vaccines differ in their storage requirements. This difference in temperature requirement lies in the composition of the vaccines, stabilising agents and active ingredients.
The dream at the heart of the scientific community has become true. The manufacturing of mRNA at record pace holds the potential to advance medicine. The applications of mRNA are not just limited to mRNA vaccine technology. Stay tuned for the upcoming blog post that peek into the past and future of mRNA applications and mRNA vaccine technology.
Are you working on a mRNA application or SARS-CoV-2? Eurofins Genomics provides all you need for your research and development.
SARS-CoV-2 Research
Control plasmids for SARS-CoV-2 research
SARS-CoV-2 (COVID-19) RT-PCR assay kit
SARS-CoV-2 Testing
ARTIC NGS primer pool for SARS-CoV-2 whole genome sequencing
ARTIC SARS-CoV-2 whole genome sequencing
High quality SARS-CoV-2 full-length genome sequencing
By Tamseel Fatima
Did you like this article? Then subscribe to our Newsletter and we will keep you informed about our next blog posts. Subscribe to the Eurofins Genomics Newsletter.




2 Comments