The early beginning of DNA sequencing
The progress from the first isolation of DNA by Friedrich Mietscher in 1869 to next generation sequencing (high-throughput sequencing) was the result of continuous efforts of the science community. After Watson, Crick and Franklin discovered the structure of the DNA in 1953, many attempts were made to sequence DNA. Eventually, in 1965, Robert Holley sequenced the first tRNA, for which he was awarded the Nobel Prize in 1986. In his Nobel Prize speech, he said, “without minimizing the pleasure of receiving awards and prizes, I think it is true that the greatest satisfaction for a scientist comes from carrying a major piece of research to a successful conclusion” (Holley, 1968). In 1972, Walter Fiers was first to sequence the DNA of a complete gene (the gene encoding the coat protein of the bacteriophage MS2) by utilising RNAses to digest the virus RNA and isolate oligonucleotides, and then separating them via electrophoresis/chromatography (Declercq et al. 2019; Min Jou et al., 1972).
The breakthrough in DNA sequencing: The first generation
In parallel to Fiers achievement, Fredrick Sanger kept working on an alternative DNA sequencing method and in 1977, developed the first DNA sequencing method that utilised radiolabelled partially digested fragments called “chain termination method”. This method went on to dominate the sequencing world for the next 30 years! Frederick Sanger is considered as a giant in genomics. He was awarded the Nobel Prizes for his revolutionary work in 1980 (his second Nobel Prize; in 1958 he was awarded his first Nobel Prize for his work on the structure of insulin). He said, “Scientific research is one of the most exciting and rewarding of occupations. It is like a voyage of discovery into unknown lands, seeking not for new territory but for new knowledge. It should appeal to those with a good sense of adventure” (Berg, 2014).

In 1977, Sanger also used his method to sequence the first ever complete genome: the one of the bacteriophage PhiX174 (virus that infects E. coli). Later, it became the most popular DNA positive control in labs around the world.
Figure 1: Frederick Sanger
Also in 1977, Maxam and Gilbert introduced a method for DNA sequencing that was based on chemical modification of DNA. The method involved using chemicals that break the DNA sequence at specific bases (Gužvić, 2013; Heather and Chain, 2016). In contrast to Sanger sequencing, it did not rely on DNA polymerase. However, both Sanger sequencing and Maxam–Gilbert sequencing lacked automation and were time consuming and tiring. Nevertheless, the research community had recognised the potential of Sanger sequencing by then, and many research groups worked on automation of the process.
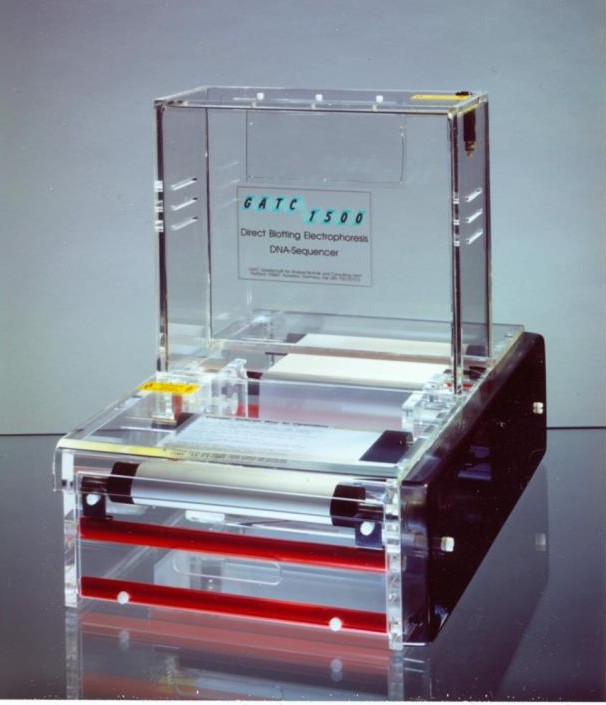

In 1984, Fritz Pohl established the first sequencing technology platform that did not rely on radioactive labelling: the GATC1500.
Figure 2: The direct blotting electrophoresis system GATC1500.
In 1987, Leroy Hood and Michael Hunkapiller who worked at Applied Biosystems, Inc. (ABI) succeeded in the automation of the Sanger sequencing process. They brought two major improvements to the method. Firstly, DNA fragments were labelled with fluorescent dyes instead of radioactive molecules. Secondly, data acquisition and analysis was made possible on the computer. The resulting instrument was named as ABI 370 (Gužvić, 2013; Hood et al., 1987).
Due to the continues innovative efforts, DNA sequencing was taken to new heights and, eventually, to the next generation.
Innovation accelerates: Start of the next generation
In 1996, Mostafa Ronaghi, Mathias Uhlen and Pȧl Nyŕen introduced a new DNA sequencing technique called pyrosequencing, and that is considered as the emergence of the second generation of DNA sequencing. This automated technology is based on the measurement of luminescence generated as a result of pyrophosphate synthesis during sequencing (sequencing-by-synthesis technology), and it could already be classified as high-throughput sequencing. Later, other biotechnology companies, hosting their own technologies appeared. In 1998, Shankar Balasubramanian and David Klenerman who founded Solexa, developed a new sequencing-by-synthesis method that utilises fluorescent dyes.
The key feature of NGS is parallelisation of a large number of reactions. This is achieved through automation and miniaturisation of the reactions.
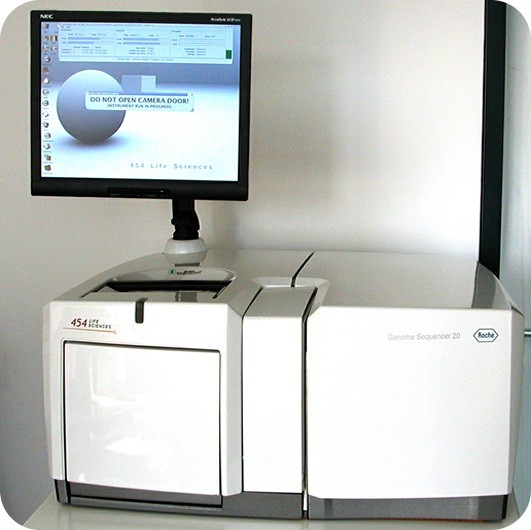

2005 was the year when Jonathan Rothberg and colleagues implemented the pyrosequencing technology in an automated system: The 454 system that was the first next generation sequencing platform to come to market.
Figure 3: Roche 454 Sequencing System.
When Illumina acquired the company Solexa in 2007, they set out to provide the most widely used NGS technology in the world and they are the NGS platform market leader to this day.
Other notable platforms that are based on different technologies are SOLiD system’s “sequencing-by-ligation” in 2007, and the Ion Torrent by Life Technologies in 2011 that uses “sequencing-by-synthesis” technology that detects hydrogen ions when new DNA is synthesised.
Enter the third generation of DNA sequencing
It is difficult to distinguish the third generation sequencing technologies from the second generation. Arguments are made that single molecule real-time (SMRT) sequencing technologies should be defined as the third generation (Heather and Chain, 2016). Pacific Biosciences, Inc. (PacBio) is the pioneer of third generation sequencing technology. In 2010, the introduction of its zero-mode waveguide (ZMW) raised the bar for DNA sequencing methods. ZMW utilises “nanoholes” that contain a single DNA polymerase. Here, the incorporation of a single nucleotide can be observed directly. Each nucleotide is labelled with a different fluorescent dye, and the signal that is emitted during incorporation is immediately recorded and read by very strong detectors attached below the ZMW.
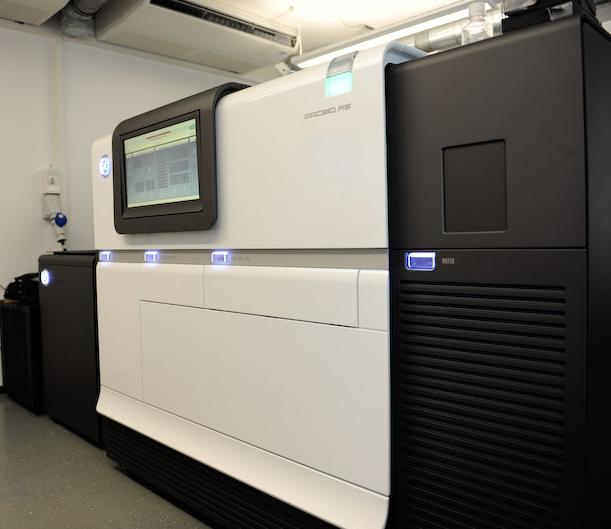

Eric Schadt, PacBio’s Chief Scientific Officer explained, ‘‘We can multiplex that process and observe this phenomenon as it happens over thousands or tens of thousands of these holes simultaneously. The DNA polymerase becomes the sequencing engine. That is a true revolution” (McCarthy, 2010).
Figure 4: PacBio RSII sequencer.
The latest versions of single-molecule sequencing systems have been drastically reduced in size. Oxford Nanopore Technologies’ systems such as GridION, MinION or Flongle are portable handheld system for RNA and DNA sequencing for reads of more than 2 Mb. The GridION was first introduced in 2012 and uses the changes in electrical conductivity that occur when DNA strands pass through biological nanopores in order to identify the nucleotide sequence (Hayden, 2012; Lu et al., 2016).
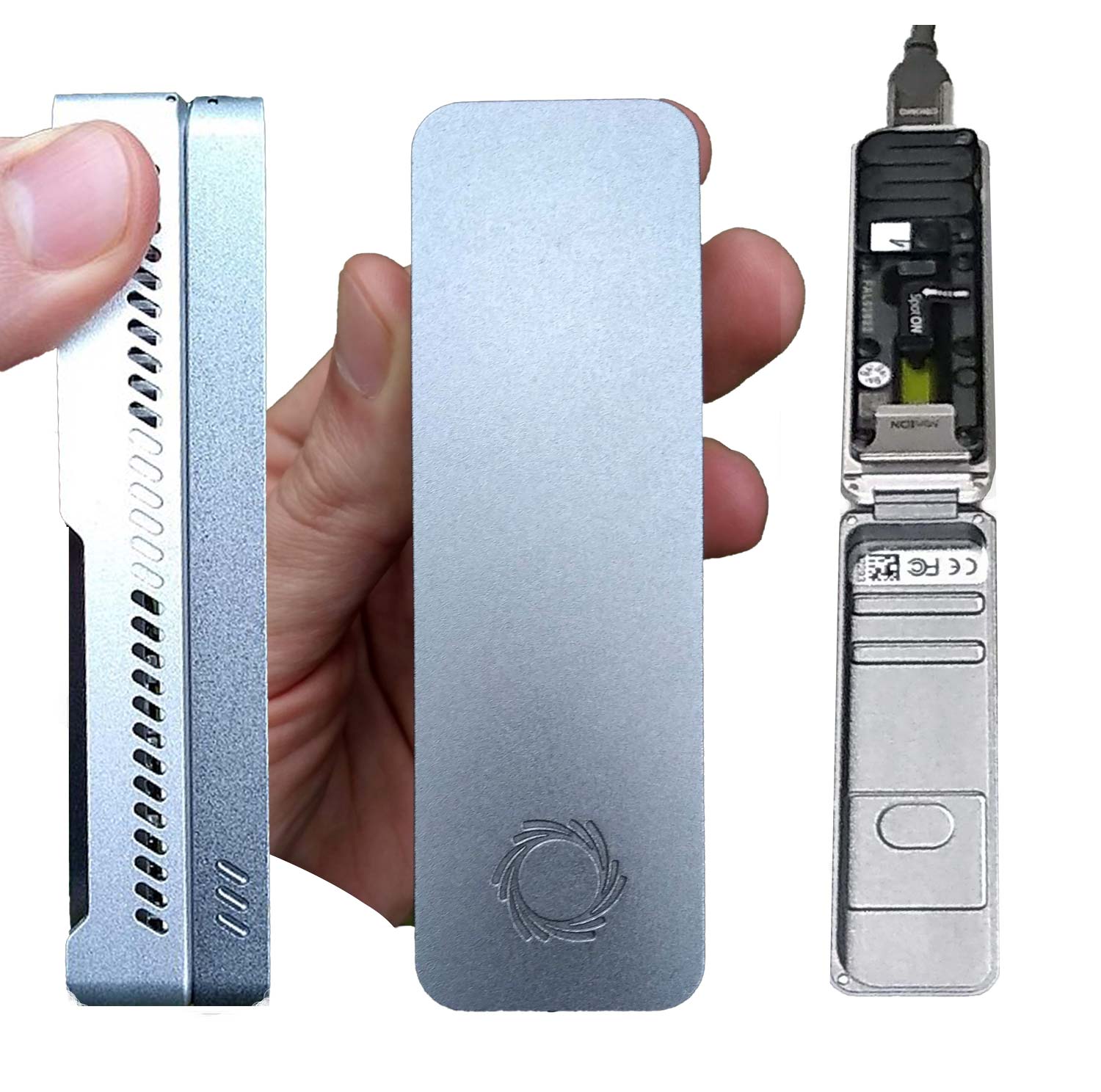

Figure 5: Image 5: Oxford Nanopore Technologies MinION.
Illumina’s NovaSeq platforms reached new hights in sequencing power for commercial platforms. Per run, the NovaSeq 6000 (S4 flow cell) generates an output of up to 3000 Gb (Illumina, 2017).
Another notable technology is Nabsys’ HD-Mapping that uses sequence-specific tags to label long DNA fragments that are then detected by nano-detectors and subsequently compiled to a map of the genome.
How does NGS work?
Presently, NGS includes various technologies that perform sequencing and gather data from multiple reactions running simultaneously. This is the reason NGS is also referred to as massive parallel sequencing. What has made NGS so powerful? NGS techniques follow the principle of Sanger sequencing, however, in Sanger sequencing, there are separate steps for sequencing, separation and detection, whereas, NGS uses array based technologies (Ilyas, 2017). Even though, there are many NGS platforms available, all of them follow the three following general steps:
- Sample/library preparation: A library is prepared by fragmenting the DNA sample and ligating it with commercially available adapter molecules. Adapter molecules act in the hybridisation of the library fragments to the matrix. Moreover, adapter molecules provide a priming site.
- Amplification and sequencing: The library is converted into single stranded molecules. Amplification, subsequently, creates clusters of DNA molecules. Each cluster acts as an individual reaction where sequencing, called run, is performed.
- Data output and analysis: At the end of the reaction, each NGS run provides a large amount of raw data. This data can be analysed by using a variety of available software.
Differences among available NGS technologies lie in the details of amplification and sequencing reaction. PacBio SMRT, on the other hand, does not involve any amplification step.
With ever-increasing throughput and decreasing costs, NGS is used for a countless number of applications in basic and applied research, biotechnology, agriculture, diagnostics, health-care, and many other fields. It may even soon be utilised as a routine diagnostic tool for diseases as part of personalised medicine. Liquid biopsy for non-invasive cancer detection for example already applies NGS in an innovative way. Exciting times ahead!
Did you like this article? Then subscribe to our Newsletter and we will keep you informed about our next blog posts. Subscribe to the Eurofins Genomics Newsletter here.
By Tamseel Fatima and Dr Andreas Ebertz
Are you unsure what NGS approach to pick for your scientific question and investigation? Try our NGS decision tree for human and mammalian samples or non-human samples to find the best fitting service.
In case the depiction of the decision tree does not work properly in Internet Explorer or Firefox, please use Google Chrome.





It’s interesting to learn that the DNA structure was discovered in the year 1953. I also never knew that the first NA sequencing method was developed in 1977. Anyhow, if I were those people who would like to discover the building blocks of the genetic code must consult with a laboratory service.