The problem of human-like-protein production
Lots of proteins for therapeutic uses are already being produced with the help of modern synthetic biology. A prominent example for this is insulin, the protein responsible for coordinating blood sugar levels in humans. Ever since the late 1970, it is synthesised within genetically modified E. coli cells. The resulting recombinant insulin is identical to its humane pendant and therefore absolutely safe to use in a therapeutic context.
As easy and accessible as transgenic protein production is, its efficiency is unfortunately subject to many biological limitations. Most human endogenous proteins undergo various post-translational modification steps which, during transgenic expression, have to be meticulously reproduced in order to prevent unwanted immune reactions once applying the produced protein in living humans. Most lab organisms used for recombinant protein production, such as E. coli, struggle with recreating such complex post-translational modification patterns.
Insulin is a lucky exception here, because unlike most of the human endogenous proteins, it barely undergoes any post-translational modification at all. This makes it very easy to be synthesised in different organisms such as E. coli. The small bacteria have a high reproduction rate and are easily cultivated, yet they lack the ability to reproduce eukaryotic post-translational modification patterns in sufficient detail. This creates the need for another microorganism more suitable for the production of human therapeutical proteins. This is where our project comes into play!
The solution to human-like-protein production
Introducing the new bright star in the world of lab organisms – Leishmania. These protozoic single-cell parasites can be very dangerous to their host animals, but they hold the key to a more efficient production of complex human proteins. The post translational modifications that proteins in Leishmania undergo are highly similar to those in human cells. This creates the possibility to produce human-like modified proteins in Leishmania. And much like E. coli, they are easy to cultivate and grow fast and in large quantities. Even the expression of large and complexly structured proteins can be efficiently realised in Leishmania, something that other lab organisms often struggle with.
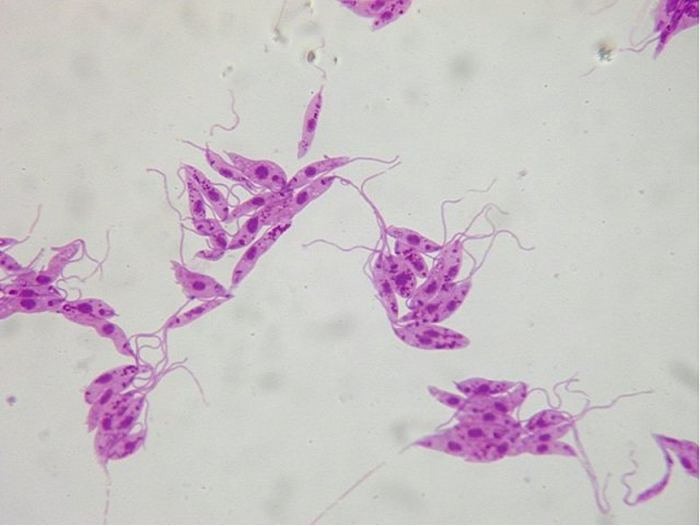
Figure 1: Leishmania in close-up. These little fellas were isolated from an infected animal’s blood, stained and can now be kept on medium in the lab and reproduce.
One of the most frequently used strains is Leishmania tarentolae which, like all Leishmania strains used in research, is non-pathogenic to humans and therefore completely safe to use in the lab.
So far so good. But as easy it may have sounded up to now, the introduction of a new metabolic pathway into an organism takes a lot of time and work. To solve this problem, scientists have developed a method called modular cloning. This method developed by Ernst Weber et al. uses a Building-Block-System that allows for easy assembly of different genetic parts resulting in a fully functional gene.
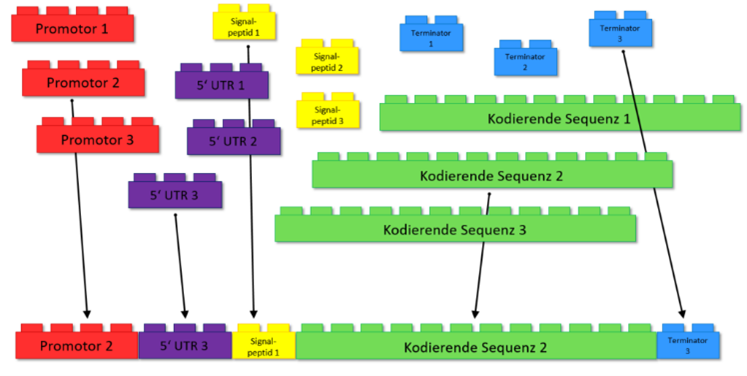
Figure 2: Lego for biologists. The different parts of a gene can be assembled like Lego bricks.
Modular cloning is based on the Golden Gate Assembly method, which was established by Marillonet et al. in 2008.
This system makes use of Type IIS restriction enzymes which, unlike most other restriction enzymes, cleave DNA not within, but in a specific location outside of their recognition site. This cleavage results in so called scarless overhangs, which do not include the recognition site and therefore, once assembled, cannot be cleaved again. The advantage: Cleaving of the module and assembly into the designated vector can take place simultaneously in one reaction.
The second advantage of this system is that since Type IIS restriction enzymes always cut at the exact same distance from the recognition site, you can choose what nucleotides the remaining overhang should contain. This allows you to specify the overhangs of the modules in a way that makes them fit to one another in the correct order and assemble all of them in this exact order, again, in just one reaction.
When establishing this system for an organism, the nucleotide overhangs functioning as the transitions between modules are standardised. This allows you to select individual genetic parts from your finished module library and combine them at will, knowing that they will always assemble in the correct order.
Our project on human-like-protein production
Our goal is to introduce the modular cloning system into Leishmania tarentolae. This requires us to create the genetic building blocks specifically tailored to the transcription and translation machinery found in Leishmania. Once we have this down, we want to create our own little library of genetic parts which can then be used by other research groups in order to express proteins in Leishmania tarentolae as desired. As a first model contestant we will try to express the receptor binding domain of the SARS-CoV-2- spike protein and are hoping to hereby showcase the current medical relevance of our project.
The team
We are a team of 9 students of the Biology Department at the Technical University of Kaiserslautern. We are mainly being supported by the AG Molecular Biotechnology of Professor Schroda and the AG Biochemistry of Professor Deponte.
Together, we are competing in iGEM, an international competition for synthetic biology. Every year more than 300 teams from all over the world participate with innovative ideas and approaches. In the end of October, they all come together in a big final event called Giant Jamboree where they can present their projects in front of the large, international iGEM community.
Besides the lab work we also have a few other projects regarding public relations. One of our current projects is a Science Slam that we are organizing together with our Cooperation Partner iGEM Bonn. It will take place on June 18th online via twitch. We’d love to meet you there! Check out our social media for more detailed information on this and many other upcoming projects!
If you want to know more about the project of “iGEM TU Kaiserslautern 2021” visit their website iGEM 2021 – TU KAISERSLAUTERN (uni-kl.de) or their social media (Facebook @igem.kaiserslautern; Instagram @igem.kaiserslautern; twitter @iGEM_TUKL) or contact them via email igem@bio-uni-kl.de.
By Nicolas Bayer and Julia Spänle
Did you like this article? Then subscribe to our Newsletter and we will keep you informed about our next blog posts. Subscribe to the Eurofins Genomics Newsletter.